Artificial intelligence in medicine
Modern analytical techniques provide physicians with an ever-increasing amount of information, the analysis of which is becoming increasingly difficult without the help of computers. This flood of data has led to intense change in the field of data analysis over the past two decades. While the original way of programming was to teach the computer to solve problems by means of well-defined rules, nowadays the models developed are capable of learning on their own and are thus getting closer and closer to artificial intelligence.
Instead of giving the computer the rules, sample data is collected (e.g. images, texts, audio) from which the algorithm (=computer program) independently selects and extracts the relevant information and creates its own rules. Such algorithms are often based on the functioning of the human brain and are therefore called "neural networks".
Artificial Intelligence in Cytomorphology
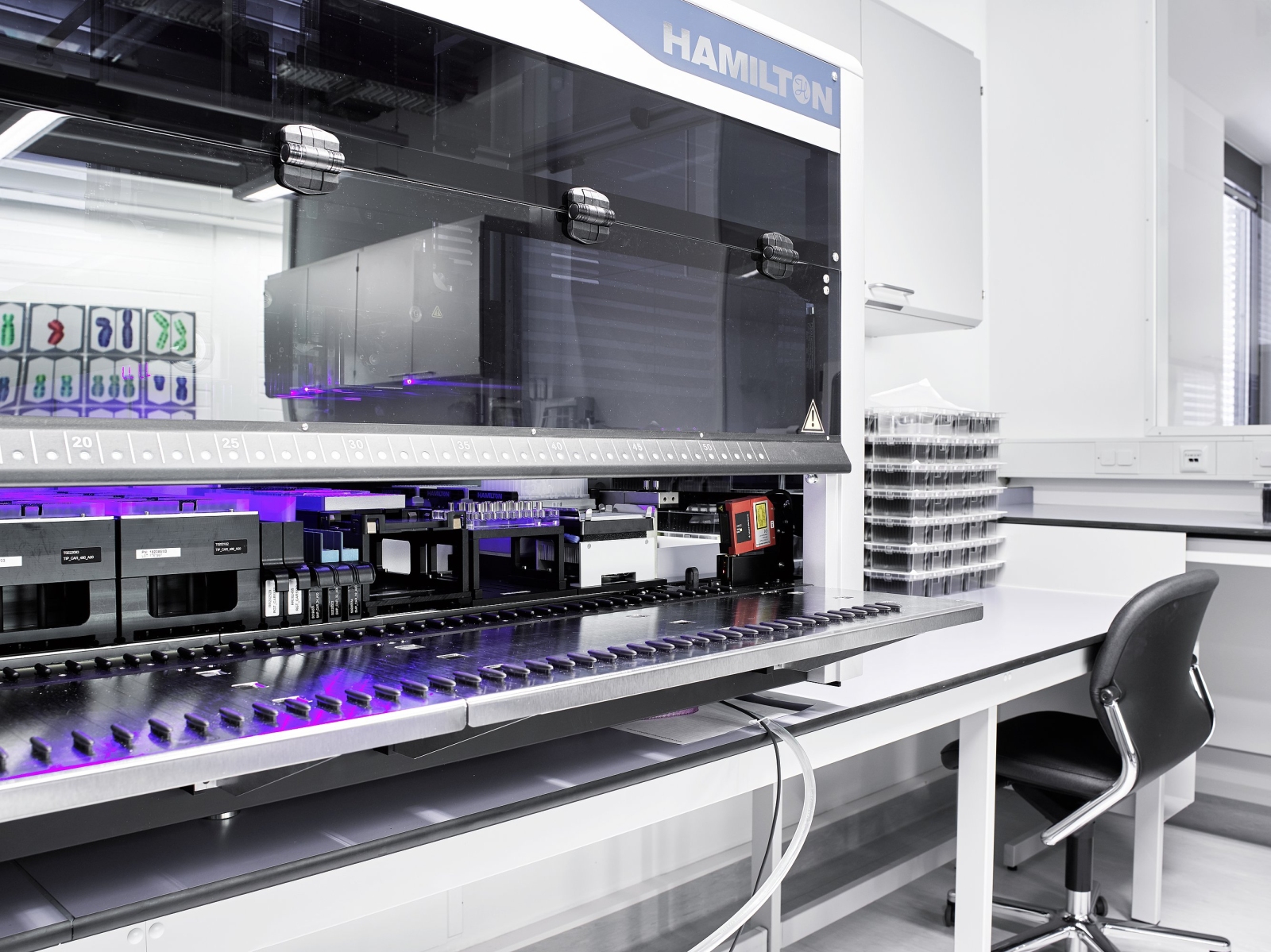
In medicine, there are several areas in which artificial intelligence is or can be applied. Particularly advanced is the use of neural networks in image recognition. Following the WHO guidelines, the diagnosis of leukemia is still strongly influenced by cytomorphology. Cytomorphology focuses on the assessment of blood and bone marrow smears to describe and differentiate malignant and healthy cells. Here, the morphologist looks for abnormal patterns in appearance and number of different cell types, which are then classified using established guidelines.
However, the quality of the result depends heavily on the experience of the morphologist, and even with experienced and trained hematopathologists, the reproducibility is only 75 to 90%. In addition, manual evaluation can be quite tedious and time-consuming, limiting the number of cells that can be processed per sample and sample throughput in general. However, advances in digital microscopic imaging and various machine learning techniques have made automated image processing and classification possible. To standardize the process of differentiating peripheral blood cells, we established a workflow to automatically acquire and digitize microscopic images of blood smears and, in collaboration with AWS, trained an machine learning (ML) model that identifies 21 predefined classes of different cell types. The blood smears are initially scanned at 10x magnification to define the relevant area and then images of individual cells are generated by a high resolution 40x scan. These images are then fed to the ML model, which returns the class (= cell type) with the highest probability for each image. In a first interim analysis, the comparison of the results for cell differentiation between the experts and the ML model showed a high agreement, so we are confident that the method can soon be used in routine to support hematologists. In another project, we are working in collaboration with the Institute for Artificial Intelligence in Healthcare (Helmholtz Zentrum, Munich) on automated analysis of bone marrow smears, and the initial results are extremely promising.
Here, as with all ML-based algorithms, the more data available, the more accurate the prediction. The accuracy of a morphologist also increases with his experience - the more time he has spent in front of a microscope and the more extensive the range of smears viewed, the more accurate and rapid his assessment becomes.
Artificial intelligence in Cytogenetics
Similar methods can be applied in all areas which are primarily based on the analysis of image files. The greatest success to date in the use of artificial intelligence at MLL has been achieved in cytogenetics. Here, an ML-based system has already been used since November 2019 to automate various steps in chromosome analysis. Chromosome analysis is about obtaining patient-specific information by classifying chromosomes and detecting any chromosomal aberrations. Here, the chromosomes are classified based on size and banding pattern and displayed in a karyogram. However, the generation of such a karyogram is a very time-consuming and complex process. For example, the chromosomes in the recorded metaphases must first be carefully separated from each other before they can be assigned to their place in the karyogram. The automatic separation of individual chromosomes is not a trivial task, as overlaps also occur from time to time. However, since February 2021, an algorithm for automatic chromosome separation has been used in cytogenetics at MLL, which only requires conditional manual support/correction. Already since November 2019, the use of a trained and optimized neural network allows the automatic classification of individual chromosomes and the generation of karyograms for patients without cytogenetic alterations. In the summer of 2021, this algorithm was further optimized so that all recorded metaphases per patient are now analyzed simultaneously. This has further increased the number of cases reported within 7 days. Further improvements have led to the fact that numerical aberrations (gain or loss of whole chromosomes) are now also reliably classified. Structural aberrations (e.g. translocations, inversions, etc) are more challenging, but during automatic classification chromosomes that are clearly different from normal chromosomes are sorted out for manual classification, saving time even for aberrant karyotypes.
Artificial intelligence in Immunophenotyping
ML-based models are also used at MLL in immunophenotyping, where malignant cells are distinguished from healthy cells based on their antigen expression pattern using flow cytometry. The individual cell types are characterized by the expression of specific antigen combinations. Diagnosis of the various hematologic neoplasms is made by interpreting the recorded two-dimensional graphs of flow cytometry. Each analysis involves measuring thousands of cells, which greatly increases the amount of data. In collaboration with AWS, several ML-based models have been trained using the raw flow cytometry data, allowing the classification of six different subtypes of hematologic neoplasms (AML, MDS, ALL, T-NHL, B-NHL, multiple myeloma/MGUS). The models are currently being tested and evaluated in routine settings. We anticipate that the trained models will replace up to 75% of routine data analysis in Immunophenotyping in the future. Our next steps here focus on classifying additional entities, applying transfer learning to achieve universal applicability, and extending the models to also detect measurable residual disease patterns.
Artificial intelligence in Molecular Genetics
In molecular genetics, the increase in sequencing performed means that data volumes are growing and manual interpretation of the data is becoming increasingly difficult. While previously limited to the study of individual genes, high-throughput sequencing methods allow the simultaneous study of the entire genome (WGS) and/or transcriptome (RNA-Seq). The goal of these methods is not only to analyze gene-specific changes and/or overexpression in a high-throughput manner, but rather to uncover underlying regulatory mechanisms and identify recurring genetic patterns. For example, are there certain combinations of genetic alterations that characterize the clinical picture of a particular type of leukemia? Several molecular markers are already known to distinguish the different subtypes of leukemia, but knowledge is still limited. The immense amounts of data make manual sifting through genomic data impossible, and since you don't know what you're looking for, you can't tell a computer how to find it. For this reason, machine learning methods are used, which learn independently from the data and extract relevant information. In principle, this approach pursues two goals: on the one hand, one wants an automatic classification of unknown samples and, on the other hand, one wants to gain further insights into the fundamentals of the various diseases. In order for this to work, the algorithm, which is often a neural network, is trained on genomic data of the various subtypes and its performance is evaluated. This is a highly iterative process to find the optimal setting of parameters that guarantee the best performance and thus the most accurate classification. Even though the genome of different people is 99.9% identical, they differ in a large number of polymorphisms. To prevent these individual differences from negatively affecting the classifier's performance, large amounts of training data are needed to cover the diversity that occurs and guarantee accurate estimates. Since the algorithm searches itself for the features for the individual subtypes, it stands to reason that this will also allow new correlations and associations to be found that may help to better understand the molecular basis. This, together with the already known features of routine diagnostics, should make improved diagnosis and prognosis assessment possible.