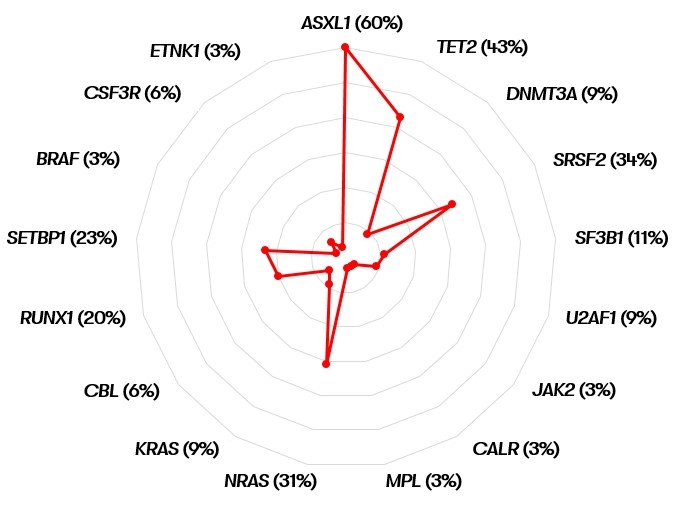
Myelodysplastic/myeloproliferative neoplasm with neutrophilia (MDS/MPN-N)
- Method:
- Anticoagulant:
- Recommendation:
- Method:Cytomorphology
- Anticoagulant:EDTA
- Recommendation:obligatory
- Method:Immunophenotyping
- Anticoagulant:EDTA or Heparin
- Recommendation:facultative
- Method:Chromosome analysis
- Anticoagulant:Heparin
- Recommendation:obligatory
- Method:FISH
- Anticoagulant:
- Recommendation:no
- Method:Molecular genetics
- Anticoagulant:EDTA or Heparin
- Recommendation:obligatory
Based on the current guidelines and the current state of research, there are different diagnostic recommendations for patients with MDS/MPN-N (myelodysplastic /myeloproliferative neoplasm with neutrophilia). We have summarized the most important information on classification and diagnostic methods at MLL.
MDS/MPN-N: Classification
Myelodysplastic/myeloproliferative neoplasm with neutrophilia (MDS/MPN-N) is a myeloid neoplasm with myelodysplastic and myeloproliferative features characterized by persistent neutrophilia in peripheral blood and neutrophilic left-shift (Khoury et al. 2022). The name MDS/MPN-N was introduced in 2022 with the appearance of the new WHO classification, replacing in it the designation of the entity as atypical chronic myeloid leukemia (aCML), BCR::ABL1-negative. The change in name emphasizes the myelodysplastic/myeloproliferative nature of the disease and is intended to avoid confusion with chronic myeloid leukemia (CML) (Khoury et al. 2022). In the International Consensus Classification (ICC), also published in 2022, the entity is referred to as atypical chronic myeloid leukemia (Arber et al. 2022).
Table 1: Diagnostic criteria in MDS/MPN-N (Khoury et al. 2022)
|
Essential:
|
|
Desirable:
|
* Diagnosis of MDS/MPN-N requires exclusion of a BCR::ABL1 fusion gene, which requires careful evaluation to exclude cryptic rearrangements and/or alternate BCR::ABL1 transcripts using available methods (e.g., cytogenetics, fluorescence in situ hybridization, or PCR-based analyses).
** Mutations in these genes are uncommon in MDS/MPN-N and should prompt morphologic examination to rule out alternative diagnoses.
MDS/MPN-N: Diagnostic methods and their relevance
MDS/MPN-N: Prognosis
The median survival of MDS/MPN-N patients is 14-29 months (Khoury et al. 2022). However, because it is a rare disease, there is no uniform consensus on risk stratification (Patnaik & Tefferi 2023). As a prognostic-negative parameter in MDS/MPN-N, a leukocyte count of >50x109 /L has been described in several studies (Onida et al. 2002, Breccia et al. 2006, Wang et al. 2014, Khoury et al. 2022). In individual of these studies, age >65 years, female gender, and hemoglobin level <10 g/dL were also prognostically unfavorable. In addition, a SETBP1 mutation was shown to have a negative impact (Piazza et al. 2013, Khoury et al. 2022, Patnaik & Tefferi 2023). 30-40% of patients show transformation to AML (Wang et al. 2014).
MDS/MPN-N: Recommendation
It should be noted that according to WHO 2022, when a CSF3R mutation is detected, CNL should be excluded morphologically as part of the differential diagnoses. Similarly, if a JAK2, CALR or MPL mutation is detected, an accelerated phase of MPN should be excluded based on history.
In contrast, according to WHO 2022, the presence of a SETBP1 or ETNK1 mutation supports the diagnosis of MDS/MPN-N (Khoury et al. 2022).
Status: June 2023